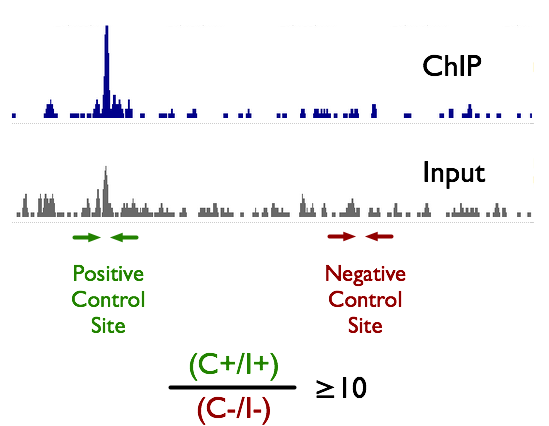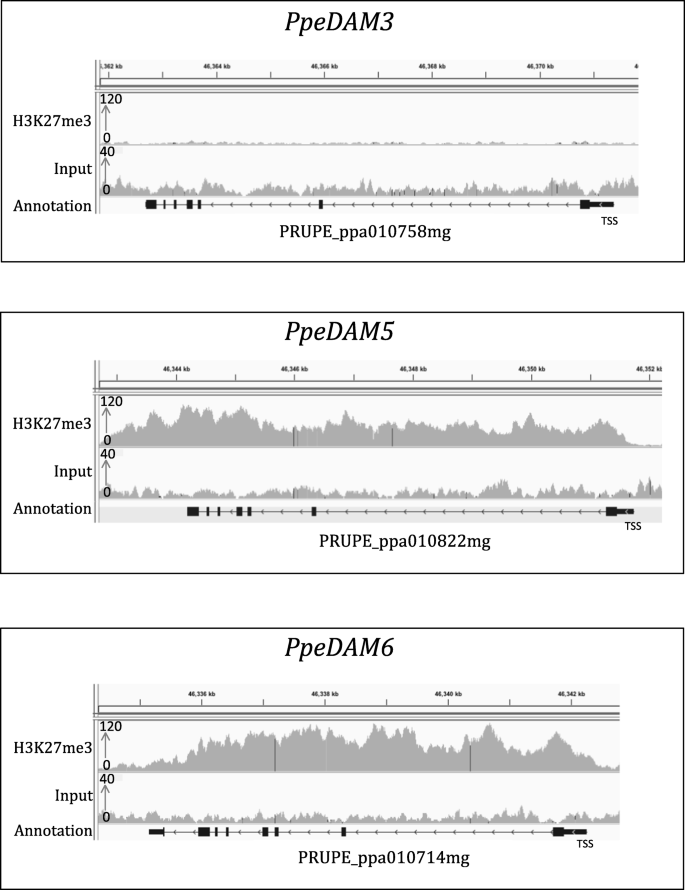
ChIP-seq and RNA-seq for complex and low-abundance tree buds reveal chromatin and expression co-dynamics during sweet cherry bud dormancy | SpringerLink

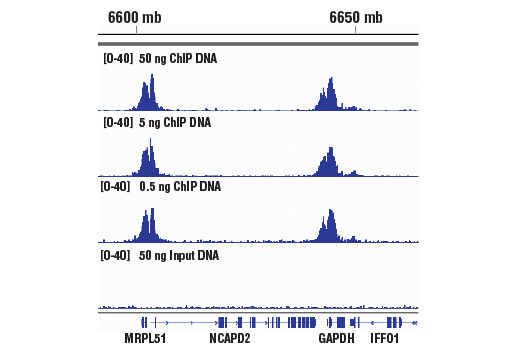
TAF-ChIP: an ultra-low input approach for genome-wide chromatin immunoprecipitation assay | Life Science Alliance
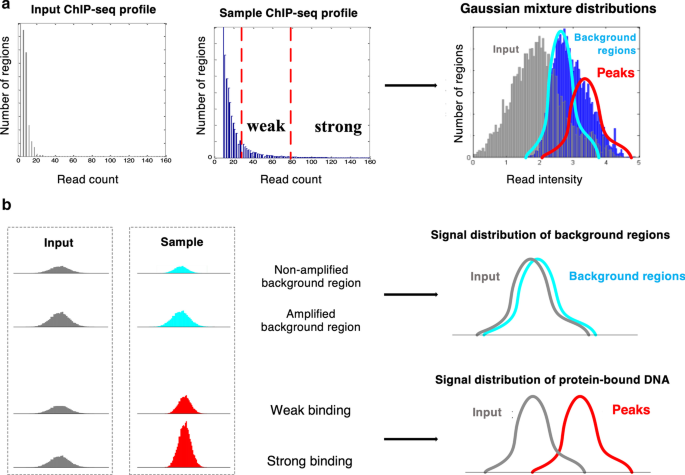
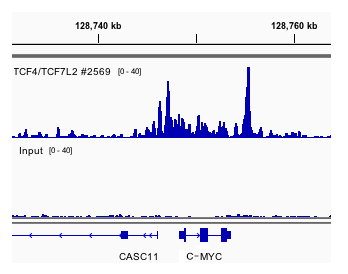
ChIP-BIT2: a software tool to detect weak binding events using a Bayesian integration approach | BMC Bioinformatics | Full Text

TAF-ChIP: an ultra-low input approach for genome-wide chromatin immunoprecipitation assay | Life Science Alliance
ChIP-seq and In Vivo Transcriptome Analyses of the Aspergillus fumigatus SREBP SrbA Reveals a New Regulator of the Fungal Hypoxia Response and Virulence | PLOS Pathogens
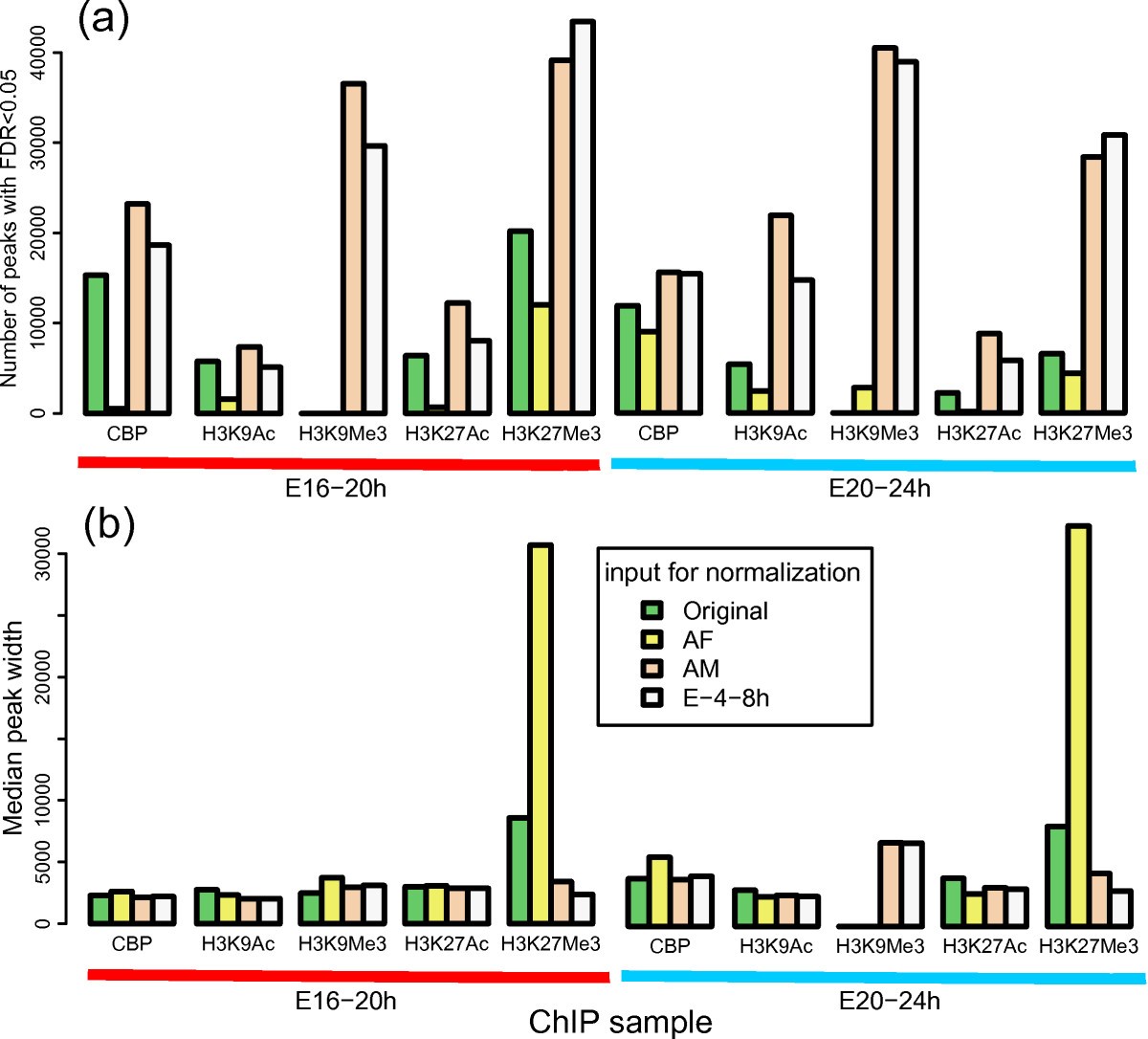
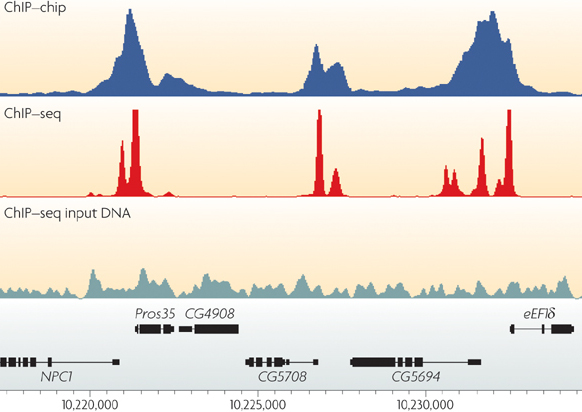
ChIP-chip versus ChIP-seq: Lessons for experimental design and data analysis | BMC Genomics | Full Text
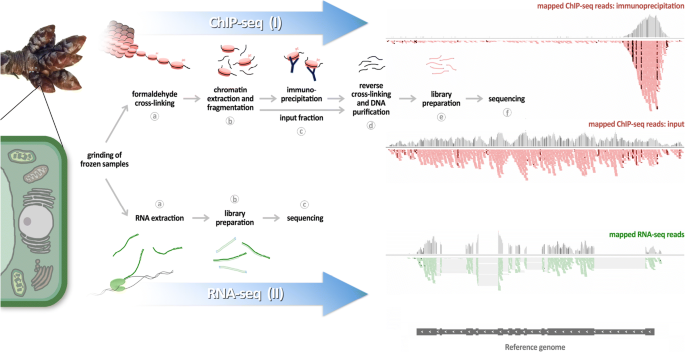
ChIP-seq and RNA-seq for complex and low-abundance tree buds reveal chromatin and expression co-dynamics during sweet cherry bud dormancy | SpringerLink

ChIP-seq profiles of AGO2 in S2 and S3 cells at BX-C. AGO2 ChIP-seq... | Download Scientific Diagram